An Introduction to ISH (in situ hybridization)
ISH (in situ hybridization) is a technique used for detecting specific DNA and RNA sequences in a tissue sample by using a labeled RNA or DNA probe. The labeled probe is detected using an antibody. It is an important method used for scientific research in clinical settings. It is also used as a diagnostic tool in understanding gene mapping, cytogenetics, gene expression, prenatal diagnosis and development. The detection of a specific nucleotide sequence is accomplished using a complementary strand that is hybridized with a probe. The hybrid is visualized using autoradiography for probes that are radioactively tagged or by development of histochemical chromogen for non-isotope labeled probes. The advantage of using ISH helps researchers study the distribution of nucleotides to understand the specific gene sequence of a target protein or gene.
Two Main Methods of ISH
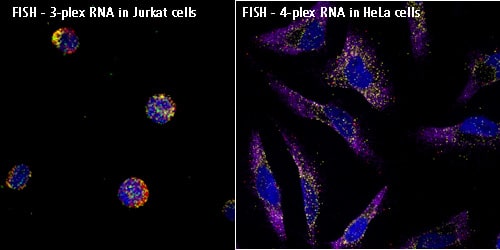
The two main methods of in situ hybridization used to visualize the RNA and DNA are CISH (Chromogenic in situ hybridization) and FISH (Fluorescent in situ hybridization). Both methods use specific nucleic acid probes which are hybridized with the tissue sample and the visualization of the product is done using either bright field microscopy or fluorescent microscopy. The tissue preparation and the methods to add the nucleotide probes are similar in both FISH and CISH but differ in the method of probe detection. The probes used in this technique ranges from locus specific probes, centromere repeats, telomere repeats to whole chromosomes.
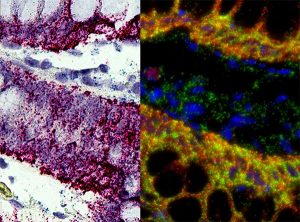
RNA in situ hybridization in FFPE samples.
ISH: In Situ Hybridization Protocol
Tissue Preparation
- Tissues used in this procedure are either frozen or paraffin embedded tissues though frozen tissues preserve the nucleic acids better. Formalin fixed tissues are also used.
- Tissues are cut .4 to .5 µm thick and loaded on slides.
- The tissues are digested with proteinase K to improve hybridization signal. It removes the proteins around the target nucleotide sequence.
- The sample is thoroughly washed with xylene and ethanol.

Image: Frozen Tissue Array
Addition of Probe
- The types of probes used are cDNA, cRNA and synthetic oligonucleotides. The RNA probe hybridizes with corresponding mRNA and DNA probe with corresponding cDNA.
- Selection of probes depends on the sensitivity and specificity of the probe, the ease of usage.
- The probes are around 40 - 1000 bases long.
- The probe is labeled either with an isotope or non-isotope.
- Denaturation happens at 95° Celsius.
Hybridization
- Dextran sulphate is added to increase the hydration in the sample for increased hybridization rate.
- Formamide and DTT (dithiothreitol) are added to allow for hybridization at lower temperatures.
- SSC (NaCl + Sodium citrate) is added to reduce the electrostatic attraction between strands.
- EDTA is added to stabilize the DNA by removing the divalent cations from the solution.
- Hybridization happens at 37° Celsius.
- The slides are washed at 70° Celsius to remove excess unbound probes and loosely bound probes.
Amplification of the Signal
- The probes are attached directly to the fluorescent molecule such as biotin, fluorescein and digoxigenin or indirectly by binding conjugated antibodies with alkaline phosphatase.
- In TSA (tyramide signal amplification) method, the antibody is conjugated with (HRP) horseradish peroxidase instead of alkaline phosphatase. It uses HRP to catalyze the deposition of biotin labeled tyramide on tissue sections.
- Biotin and DIG labeled nucleotides use indirect method of detection by using anti-DIG antibody to detect the primary DIG probe.
- The fluorescent labels are detected using fluorescent microscope to examine the cells and the tissues hybridized with labeled oligonucleotides.
Two Main Methods of ISH
FISH uses different fluorescent colors and can detect multiple target sequences simultaneously allowing for direct comparisons of different RNA types. This allows for direct visualization of genetic abnormalities and gain or loss of chromosomal segments. This method also lowers the background signals and allows for greater specificity. FISH is used for live cell imaging and improves assay speed, resolution and probe targeting. Multiplex FISH allows for detection of many chromosomal aberrations in the genome in a single hybridization procedure.
CISH uses peroxidase or alkaline phosphatase labeled reporter antibodies which interacts with a hybridized DNA probe. The tissue sample is then observed with an enzymatic reaction. Genetic abnormalities and tissue morphology are simultaneously observed in this technique.
CISH’s advantages ranges from permanent staining, low cost and the ability to use light microscopes. CISH is an alternative to FISH since bright field microscopy can be used.
In situ hybridization enables multiple hybridizations to maximize tissue usage due to finite amount of tissue sample availability from biopsies and embryonic tissues. Libraries of tissue samples can also be frozen for further research. Developments are continuously being made to increase the sensitivity of the in situ hybridization technique by either amplifying the target nucleotide sequence before the procedure or after the completion of the hybridization technique.
BioChain: Ease and Convenience
BioChain's in situ hybridization kit is designed for digoxygenin-labeled probes and alkaline phosphatase detection, eliminating the need for radioactive DNA labeling. The kit includes the pre-hybridization and hybridization solutions, NBT and BCIP detection reagents, blocking solution, SSC buffer, alkaline phosphatase buffer, and an anti-Dig antibody. The in situ hybridization kit does not include reagents and enzymes for labeling probes. Any Dig labeled detection probe can be used. BioChain’s in situ hybridization kit has protocols for both frozen and formalin fixed paraffin embedded (FFPE) tissues and cells.
Features specific to the in-situ hybridization kit:
- Useful for microRNA detection
- Non-isotopic, colorimetric detection of gene expression
- RNase-free, ISH-optimized reagents
- Can be used with either oligonucleotide or ribonucleotide probes
- All lots are tested by ISH using QC-tested probes on known tissues.
- Fully optimized protocol
- Can be used on tissue section and/or tissue microarray slides
Author
BioChain Institute Inc.