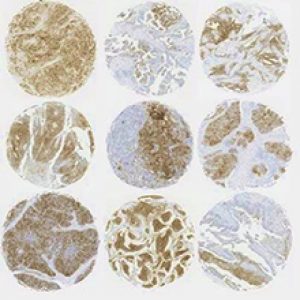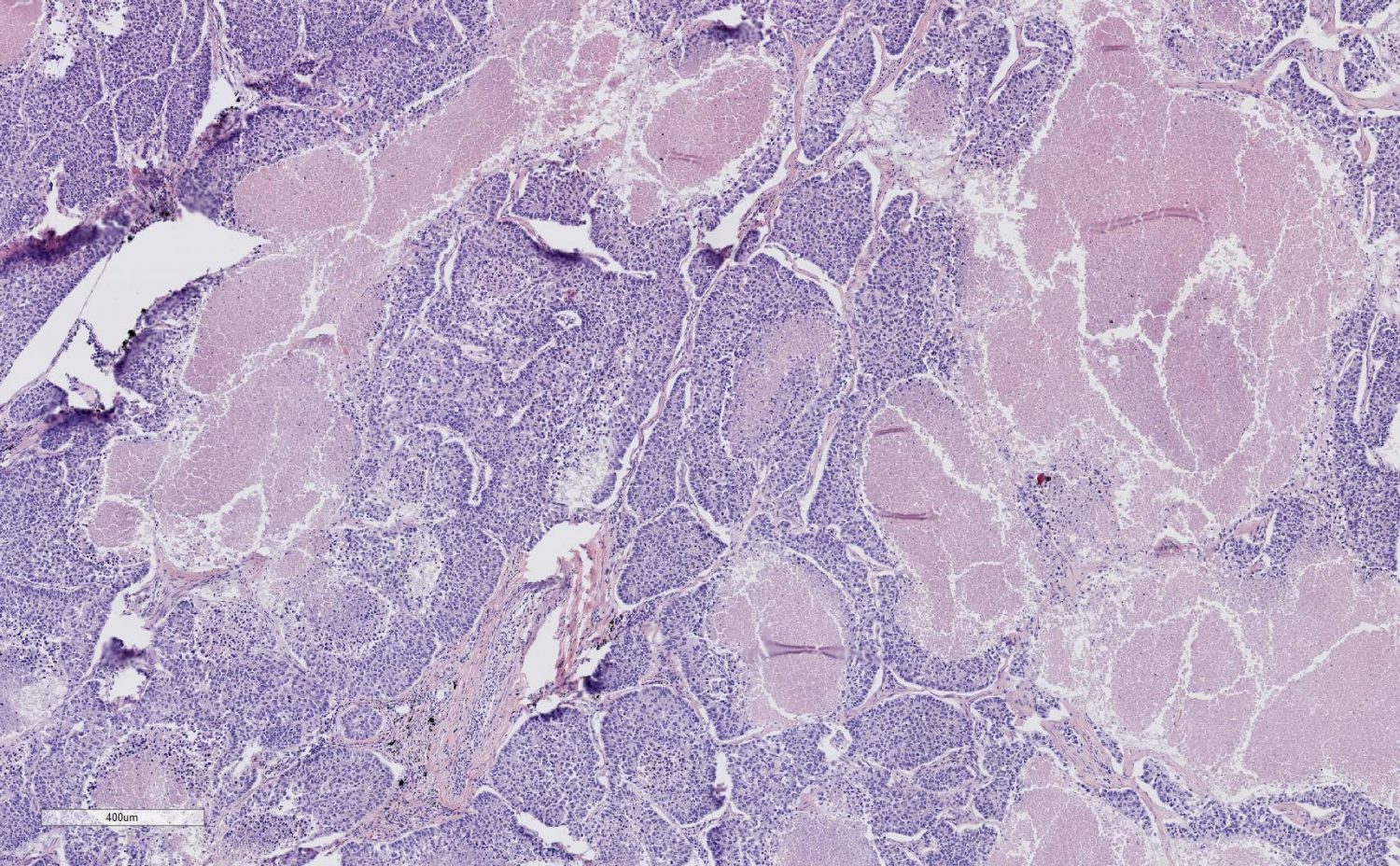
Introduction to Tissue Microarrays
A tissue microarray (TMA) is an innovative technique that can be used to analyze multiple tissue samples simultaneously. This cost-effective method has been used extensively across mediums in diagnosis, treatment, and research, proving its efficiency and versatility.
History and Methodology of Creating a Tissue Microarray
A tissue microarray arranges tiny amounts of biological matter preserved in paraffin onto a block for easy examination of hundreds, or even thousands, of samples concurrently. H. Battifora created the TMA method in the 1980s using strips of tissues wrapped around a rod of biological matter, then coated in paraffin. His “sausage” block method had many limitations, including difficulty in distinguishing individual tissue and wrapping the rod with tissue. However, once the concept was introduced, Battifora and Mehta developed a checkerboard method of arrangement, which set sections in a block pattern (Jawhar). This technique was later perfected by Wan et al, who developed the “punching” method of extraction and relocation into precise circular sections.
The final array is the result of tissue biopsies, which extract solid sample of tissue from the body. Traditional tissue biopsies, a staple in diagnoses and treatments in the medical field, usually require a substantial quantity of tissue. The TMA method reduces the amount of tissue necessary. Tissue specimen is extracted by punching into solid tissue with a circular needle that measures anywhere from 0.6 mm to 4.0 mm, depending on the amount of tissue available. The smaller these cored sections are, the more samples will be present in an array, as Figure 1 shows. The tissue is sectioned into microscopic slides, and then either formalin-fixed and embedded into a donor paraffin block for preservation, or entered into the final array directly.
When the tissue sections are ready to join the final array, they will be transferred into another block called the “recipient” block. The sections can come from multiple donor blocks. The recipient block must contain paraffin with a low melting point, and during the embedding process, care must be taken that no bubbles form within the block, or it will disrupt the array. Additionally, recipient block holes must be made immediately before the transfer occurs, otherwise the elasticity of the paraffin will cause the holes to lose their shape. Once the sections are transferred in the block, the array must be allowed to set by incubating and cooling it, making the sections firm and easier to handle. Finally, the entire array is stained, such as with immunoflourescent staining using hematoxylin and eosin, for optimal representation.
The array is organized such that X-Y coordinates are used to keep track of the tissues. Often, a spreadsheet is used to record demographic and outcome information of individual tissue sections. For this reason, recipient blocks are marked, usually at the top left, to identify the orientation of the array. Different methods of creating TMAs are demonstrated in Figure 2.
The Uses of Tissue Microarray
Tissue microarrays were originally presented in the 1980s by Battifora to test antibodies on multiple sections simultaneously. The uses of a TMA, having expanded rapidly since its popularization, are now varied in settings from research to medical services. Certain hospitals boast entire facilities dedicated only to tissue microarrays. The utilization of a TMA ranges from clinical application to laboratory research, with a substantial impact in the field of oncology. Further clinical application has been expanding in neurodegenerative, cardiac, and placental disease (Jawhar).
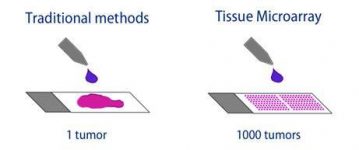
Figure 3: Traditional Biopsy vs Tissue Microarray
One of the most significant uses of a TMA is for the augmentation of rare tissue. Due to the circular precise sections with limited thickness, a single standard histological sample can yield 50-500 sections to be used in an array. Once set in the recipient block, the microarray can be further cut into 2-3 micrometer sections, doubling or tripling the number of sections. TMAs can thus produce tens of thousands of tissue disks from one sample. Since a primary benefit of a TMA is taking full potential advantage of scarce tissue material, it is possible to extract some sections for an array and return the original tissue block without significant damage. Figure 3 visualizes this process. This ensures multiple fields, such as research and diagnosis, can use the material at the same time.
TMAs are Divided into 5 Applications: Predictive, Control, Validation, Prognostic, & Progression.
With large amounts of sections available, all of them can be analyzed simultaneously. Tissues from multiple points of origin can be tested under the exact circumstance in a microarray, such as experiments that require precise uniformity in factors such as temperature, incubation time, chemical exposure, etc. Considering that each individual piece of tissue must be stained independently, variances are minimized in a control tissue microarray. Furthermore, due to entire cohorts being analyzed together, the amount of resource volume and time required is decreased, resulting in increased cost-efficiency. As a result, control tissue microarrays contribute to increased statistical quality control by providing extra samples to minimize experimental drift, a drastic improvement from using only a single tissue sample to control for quality.
As the original usage of arrays describes, predictive tissue microarrays are used in the testing of drugs, such as antibodies, to calculate responses. Predictive microarrays are invaluable in identifying drug-resistant and other therapeutic markers in genes. Other possibilities include examining how to obtain optimal staining and providing a spectrum of tissue for testing. Validation tissue microarrays corroborate biomarkers found in cDNA, such as in a study by Bubendorf et al. , checking that a certain gene expression was correlated with hormone refractory prostate cancer. The results found a 100% match.
TMAs have been irreplaceable in oncology. Through the identification of cancer-associated genetic markers and molecular profiling of tumors, microarrays assist in the diagnosis and prognosis of cancer. Using multiple tissues in one block, called multitumor arrays, can help identify novel markers and hone in on specific treatment options. In the treatment process itself, sections can be presented in tumor progression tissue microarrays, which track the expression of certain genes or proteins and determine the evolution of a cancer. Finally, prognostic tissue microarrays use staining techniques to predict the ultimate conclusion of a tumor.
Disadvantages of a Tissue Microarray
While TMAs have absolutely proved their worth, there are certain drawbacks. One such limitation is that due to the minute size of tissue section, the results may not be indicative of the entire sample. Tumors are well known to comprise of heterogeneous tissue, which expresses unique proteins in different areas. A TMA consisting of sections from that tumor may not indicate all of its genetic markers. However, a recent study by Parker et al. has shown that in a comparison for specific estrogen receptors in breast cancer, the tissue microarray analysis displayed 96% of the same markers as the whole tissue analysis. This pattern has repeated with similar consequences in multiple studies, implying that this disadvantage is not a statistical impediment. One significant disease tissue microarrays may not correlate with is heptatocellular carcinoma, a liver cancer that often presents as a heterogeneous tumor.
The Future of Tissue Microarrays
Due to the varied vital uses of TMAs, there are constant improvements in the field, including increased quantity and quality, as well as reducing the amount of time, effort, and money needed. One such advancement is amplifying the amount of tissue available for use. While current technology tends to allow around 600 sections per array, new methods are being developed that may allow up to 2000 sections of a single histological sample. This includes slicing recipient blocks micrometers thick, and taking full advantage of even finer needles to extract smaller cores.
The technique for creating a TMA is also rapidly moving toward a fully automated process, reducing time and human effort required. Automated tissue arrayer machines can work with multiple tissue blocks simultaneously. They are responsible for the coring, transfer, and recording of each section’s information. Additionally, strides have been made to convert glass slides into digital images that can be stored until necessary, as well as studied by algorithms that track certain patterns in the staining and physical features of the tissue.
Several universities, such as Stanford, are making software programs that would make it easier to obtain quality analysis and keep track of high-density microarrays through automatic digital imaging. These virtual slides are predicted to spot significant biomarkers. They can be shared virtually anywhere in the world through computers, creating a pool of specimens that can be used for long-term educational purposes. Examples of automatic imaging are demonstrated in Figure 4.
Advancements have been made in the field of TMAs for non-traditional tissue microarray platforms. Among these are using frozen tissue for an array, which has been known to affect extraction of specimens, quality of RNA present, and cause distortion in tissue morphology. Another such example is cell line microarrays, which distinguishes physical structures in a cell and could be useful to compare naturally grown cells with suspension grown cells.
It is clear that TMAs are fundamental in their function through both high-quality molecular analysis and convenience. Tissue microarrays are only just now finding their stride and will soon be an irreplaceable part of both clinical application and further research. Despite their rapid assessment of multi-origin cell blocks, TMAs need to continue evolving to keep up with the pace of automation that dominates biotechnical research.
BioChain: Leading in Samples
BioChain offers multiple prepared TMAs, as well as tissue samples that can be used in devising TMAs.
The Pancreas Tumor Tissue Array offers 102 samples of various tissues, from malignant to benign tumors, and even including normal tissue samples as a control. The TMA is particularly useful in research, such as for identifying biomarkers in the tumors and testing the effectiveness of drug treatments. BioChain offers more tissue arrays available here.

BioChain also provides human tissue panels fixed in formalin with demographic information per tissue. These are available in multi-tissue samples, or tissue specific to organs such as brain, breast, lung, etc. Panels can even be arrayed by disease, such as Alzheimer’s disease. For a complete list of BioChain’s paraffin tissue sections, search and specify here.
References
- Bubendorf et al., Hormone therapy failure in human prostate cancer: analysis by complementary DNA and tissue microarrays. Journal of the National Cancer Institution. 1999 Oct 20;91(20):1758-64. Web. Apr 2018.
- Jawhar, T. Tissue Microarray: A rapidly evolving diagnostic and research tool. Annals of Saudi Medicine. 2009 Mar-Apr; 29(2): 123–127. doi: 10.4103/0256-4947.51806. Web. Apr 2018.
- Parker et al., MD Assessment of Interlaboratory Variation in the Immunohistochemical Determinattion of Estrogen Receptor Status Using a Breast Cancer Tissue Microarray. American Journal of Clinical Pathology. 2002;117(5):723–728. Web. Apr 2018.
- Vogel, U. Overview on Techniques to Construct Tissue Arrays with Special Emphasis on Tissue Microarrays. Microarrays (Basel). 2014 Jun; 3(2): 103–136. Published online 2014 Apr 17. doi: 10.3390/microarrays3020103. Web. Apr 2018.
- Wan et al. A rapid and efficient method for testing immunohistochemical reactivity of monoclonal antibodies against multiple tissue samples simultaneously. Journal of Immunology Methods. 1987;103:121–129. doi: 10.1016/0022-1759(87)90249-3. Web. Apr 2018.
Author
BioChain Institute Inc.