
Table of Contents
What is Circulating Tumor DNA?
Deoxyribonucleic acid is a collection of nucleotides that carry genetic information necessary for the functioning of all living beings. DNA is primarily found in the nucleus of cells. When strands of DNA are found circulating in the body without a cell, it is referred to as cell free DNA (cfDNA). cfDNA exists in many forms in the body: cell free fetal (cffDNA), donor derived cell free (ddcfDNA), and circulating tumor DNA (ctDNA).
There are three major theories for where ctDNA originates (Cheng et al.) :
- necrotic/apoptic cancer cells
- living tumor cells
- circulating tumor cells (CTCs).
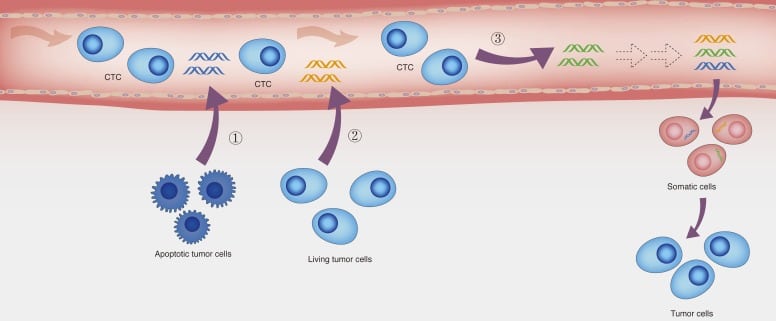
Two mechanisms have been found thus far to explain ctDNA release: passive, and active. The passive mechanism postulates that nucleic acids engulfed by macrophages from necrotic cells are released into the bloodstream. Active mechanisms may involve tumoral nucleic acids released by living cells (Calabuig et al.) . Figure 1 demonstrates potential ctDNA origins.
Levels of ctDNA increase in the blood as the amount of tumors present in the body increase, making ctDNA a vital tool in diagnosis. However, since cfDNA has a short half-life ranging from only 16 minutes to 2 hours, and ctDNA lasts about 1.5 hours, it is vital to quickly distinguish ctDNA from other cell free DNA during molecular analysis (Chang et al.) It has been found that ctDNA contains defects, such as tumor-specific mutations, copy number variations, and aberrant DNA methylations that are unique identifying factors (Warton et al.)
Function and Detection of ctDNA in Cancer
The role that ctDNA plays in cancer is still a widely researched question. One hypothesis is that cancer cells release nucleic acids with the purpose of transforming healthy cells in distant locations (Calabuig et al.) Another theorizes that ctDNA’s existence is simply a side effect of tumor cell apoptosis. While this is still unknown, one thing is certain: increased ctDNA is often an indication of increased cancer in the body, with mutations capable of being tracked directly.
Some of the most common genetic biomarkers found in ctDNA are as follows:
- KRAS – signals cells to turn proliferation on or off
- BRAF – a proto-oncogene involved in cell growth
- NRAS – regulating cell genes creating amino acids
- EGFR – creation of epidermal growth factor protein
Cell free circulating tumor DNA has been tracked in the body by researchers for decades. Beginning with the discovery of cfDNA, it was quickly discovered that ctDNA displayed characteristics of the parent tumor, such as carrying heterogenetic features and methylation patterns. It has been found in blood plasma, saliva, urine, and cerebrospinal fluid. Detecting and isolating ctDNA for real-time tracking of mutations, aiding in treatment and prognosis information is necessary.
Detection of ctDNA can be categorized using the following two methods with examples:
- PCR-based: detect DNA aberration with high sensitivity.
- Amplification-Refractory Mutation System (ARMS)
- Droplet Digital PCR (ddPCR)
- Beads, Emulsification, Amplification, Magnetics (BEAM)
- Sequencing-based: detect multiple aberrations simultaneously
- Whole genome sequencing (WGS)
- Amplicon sequencing
- Target capture sequencing
Clinical Applications and Advantages of ctDNA
Testing for ctDNA in body liquids is an invaluable part of cancer testing. The applications of ctDNA are nearly limitless, being involved from early detection of cancer to targeted treatment.
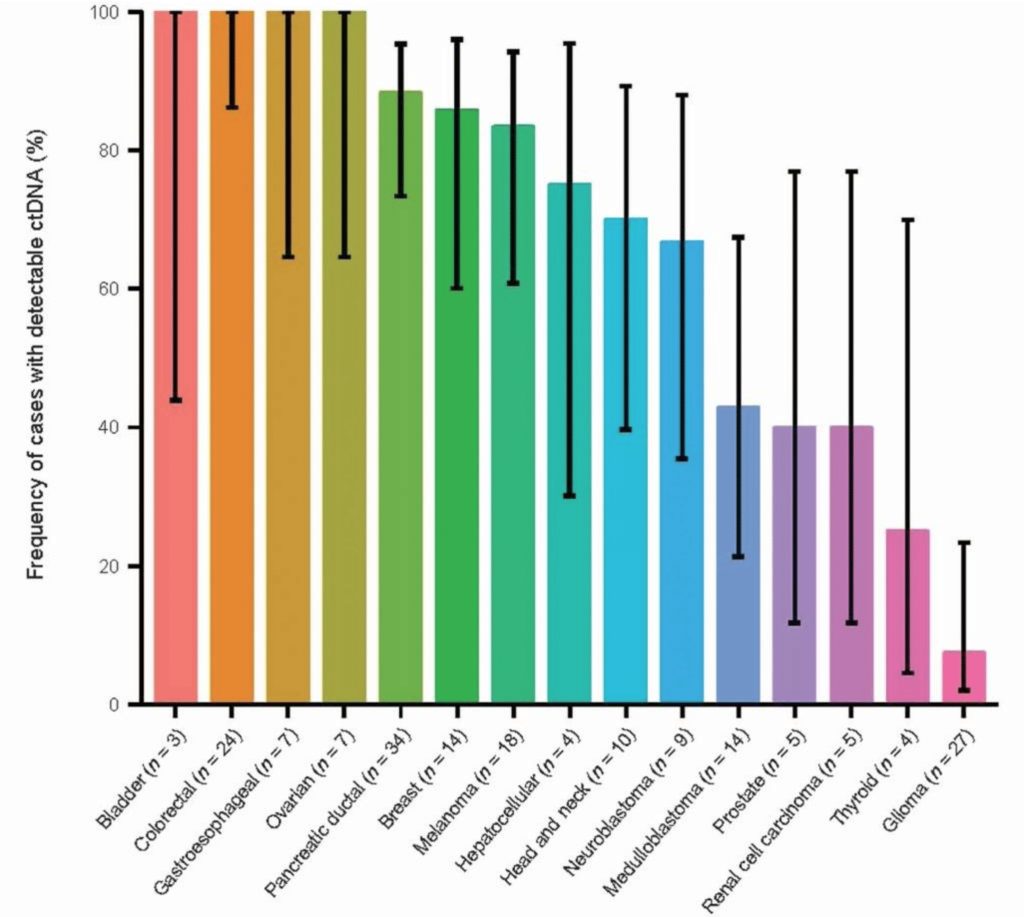
Early Diagnosis and Detection. The presence of higher levels of ctDNA is an immediate indication that malignant cancer may exist in the body. Presence of specific biomarkers helps immediately diagnose type of disease. For example, using the KRAS biomarker, Bettegowda et al. were able to find metastatic cancer in 82% of patients and localized cancer of all types in 55% of patients. Reducing morbidity and mortality can be done by early detection of cancer, leading to treatment such as surgery before the tumor metastasizes. In fact, the highest sensitivities in detecting cancer by using ctDNA are depicted in figure 2.
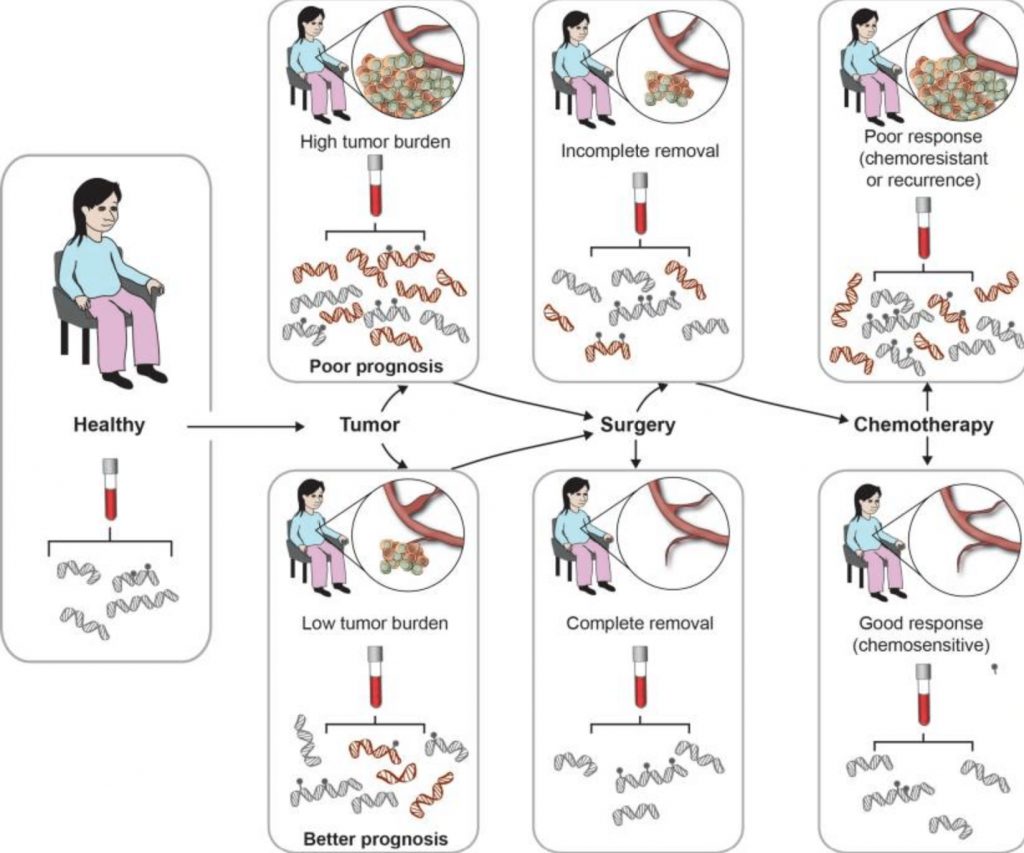
Precision Treatment (PT). ctDNA provides real-time information regarding tumors in the body. In fact, treatment-resistance mutation can also be predicted in assays, indicating the need to use different drugs or therapies. For example, KRAS codon 61 is a mutation that has been observed to be associated with acquired resistance; when this is found in ctDNA, it implies cancer cannot be treated with anti-EGFT antibodies (Loupakis, et al.) In further research, Kato et al . found a unique genotype in patients receiving radiation treatment that increased ctDNA in blood – generated only from cells destroyed by therapy. This indicated the success of treatment. PT can save patients from needless painful and ineffective treatments. Figure 3 demonstrates how PT can work with a patient.
Methylation Aberrations. In oncology, methylation refers to a change in healthy regulation of gene expression to one that indicates disease patterns. Methylation is commonly found in ctDNA, reflecting patterns found in the origin tumor. Using cancer-specific ctDNA methylation, information can be obtained on patient prognosis through measuring biomarkers (Warton et al.) This is used for both diagnosis and treatment. For example, the Epi proColon test tracks DNA methylation of the SEPT9 gene for detection of colon cancer (Epigenomics).
Monitoring Patients. Patients in remission often require close monitoring for indications of relapse. ctDNA provides a noninvasive and high-specificity method that targets biomarkers indicating the re-emergence of tumors. It also can be used to categorize patients by risk for recurrence. In a study by Douillard et al., ctDNA levels were measured in patients who underwent surgery for colorectal cancer. It was found that 100% of patients with measurable ctDNA relapsed, while patients with imperceptible amounts of ctDNA remained disease free.
Liquid Biopsy: Testing for ctDNA
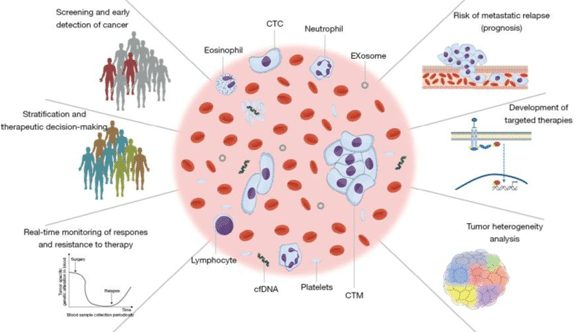
The liquid biopsy is an incredibly versatile and diverse method of isolating and detecting cfDNA in body fluids. The process uses next generation sequencing (NGS), a specific type of whole genome sequencing (WGS) which arranges and maps the DNA of an organism’s entire genome within a day (Behjati & Tarpey). Fragments of DNA are processed multiple times for an extremely detailed and accurate overview of exomes, which comprise a fraction of the genome. While the applications of liquid biopsies extend further than oncology, it is an increasingly necessary tool by cancer researchers in labs and clinics (BioChain). Some uses of the liquid biopsy are displayed in figure 4.
Traditional biopsy methods can be invasive or dangerous for patients, often qualifying as surgery, and include tissue biopsies (tissue, marrow), imaging scans (x-ray, MRI, ultrasound), and lab tests (blood, urine). Tissue biopsies may have an advantage when the tumor’s precise location is confirmed and accessible for extraction; liquid biopsies are best used as a sustainable method for identifying heterogeneous features of tumors that are unreachable by traditional methods (BioChain).
Liquid biopsies also aid in cases when tumors have metastasized. Tissue biopsies cannot feasibly reach every tumor, but ctDNA analysis can provide an excellent alternative to observing somatic mutations present in the blood.
Challenges and Limitations
While using ctDNA is emerging as a useful technique in oncology, it still has many drawbacks.
Liquid biopsies are not used as a preventative tool yet. They are not recommended for healthy people visiting the doctor. As a result, early detection of cancer is not commonplace, unless there is another persistent problem for which the biopsy was recommended.
Tracking ctDNA is also prone to false negatives and false positives. A false positive occurs when the sample indicates cancer, even if no danger is actually present – because the tumor has already been caught and eradicated by the immune system, or because it continues to grow so slowly that it will not harm the patient in his or her lifetime. False positive results can lead to overtreatment, which may cause harm to the patient.
Another consistent challenge in research has remained: certain cancers have very low or inconsistent cell death patterns, making them difficult to track (Kato et al.) Liquid biopsies can detect ctDNA, but not always the point of their origin. Extensive follow-up may be required to pinpoint an exact location.
There is no standardization in the industry yet regarding ctDNA testing. While there are many kits available for the liquid biopsy, all of the methods are different and provide results designed for various needs. Investment and standardization of the current techniques are required to make ctDNA testing accessible to all patients in a cost-effective way.
Future Research & Technological Advancements
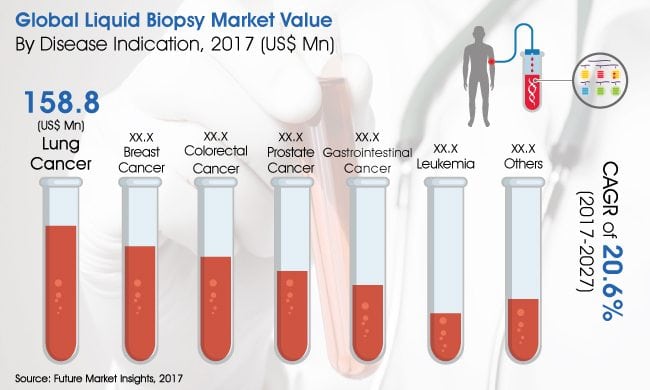
Much of the research involving ctDNA centers around liquid biopsies. As Miller et al. reported in 2016, there were approximately 15.5 million cancer survivors in the United States alone, a number rapidly increasing as technology and treatment methods improve. This demographic is at an increased risk for relapse, and liquid biopsies serve to provide patients/providers with a cheap and non-invasive alternative to regularly test for the return of cancer. The market is predicted to rapidly grow, as figure 5 shows.
While a test tracking cancer biomarkers for the pancreas is not yet ready for commercialization, it has been claimed that mutations in ctDNA shed from cancer cells and found in the blood are largely identifiable. Studies by clinical research centers have shown more than 85% of patients with advanced cancer have identifiable DNA in their blood (Bettegowda et al.) The possibility of identifying such small pieces of DNA in the blood without prior knowledge of the genetic status of the cancers demonstrates how quickly the field is advancing.
Research is also ongoing in examination of which biomarkers will be most significant in tracking cancer, as well as treatment-resistance and comorbidity. This requires the creation of universal databases that can be used to create smarter and more innovative ways of sharing data across the globe and ultimately help patients using bioinformatic tools.
BioChain’s cfPure: Quality ctDNA Extraction
BioChain is proud to offer a range of specialized products designed to extract ctDNA in blood plasma or serum. The magnetic bead-based kits can perform with less than 1 mL of a sample and can easily scale up to work with more than 10 mL of plasma or serum. In fact, BioChain’s kit recovers more cfDNA per mL of plasma when compared to competing products, as depicted in figure 6. The kits possesses convenient automation compatibility, and are rigorously quality tested for consistency in results.
Uses of the cfPure extraction kit range from NGS to PCR for DNA amplification while the scalable system allows the sample to be conserved and used only as needed. The significantly cost-effective method can be utilized for further applications, from prenatal genetic testing to early cancer diagnoses. Compared to traditional biopsy methods, BioChain’s cfDNA extraction kits can provide a more efficient method of extraction from smaller fragmented chains.

The vamPure Blood Kit also provides a method of DNA extraction from whole blood samples. The advantage of this kit is that the buffer used is free from EDTA for specific experiments. EDTA can be added separately for long-term storage. This kit also supports automation.
As well as the kits, BioChain independently provides a PCR Purification Kit - SeqPure, which increases the quality of a DNA sequence by removing enzymes, salts, buffers, and short DNA fragments. This DNA amplification kit is compatible with automation methods, saving time and costs. Leading at the forefront of biotechnological advancement, BioChain continues to couple superior products with competitive pricing to provide an exceptional clinical and research experience.
References
- Bettegowda, C., et al. (2014). Detection of Circulating Tumor DNA in Early- and Late-Stage Human Malignancies.Science Translation Medicine 6(224): 224ra24. http://doi.org/10.1126/scitranslmed.3007094
- Calabuig-Fariñas, S., Jantus-Lewintre, E., Herreros-Pomares, A., & Camps, C. (2016). Circulating tumor cells versus circulating tumor DNA in lung cancer—which one will win?Translational Lung Cancer Research,5(5), 466–482. http://doi.org/10.21037/tlcr.2016.10.02
- Cheng, F., Su, L., & Qian, C. (2016). Circulating tumor DNA: a promising biomarker in the liquid biopsy of cancer.Oncotarget,7(30), 48832–48841. http://doi.org/10.18632/oncotarget.9453
- Calabuig-Fariñas, S., Jantus-Lewintre, E., Herreros-Pomares, A., & Camps, C. (2016). Circulating tumor cells versus circulating tumor DNA in lung cancer—which one will win?Translational Lung Cancer Research,5(5), 466–482.http://doi.org/10.21037/tlcr.2016.10.02
- Douillard JY., Ostoros G., Cobo M., et al. Gefitinib treatment in EGFR mutated Caucasian NSCLC circulating-free tumor DNA as a surrogate for determination of EGFR status.J Thorac Oncol. 2014;9(9):1345–1353.
- Epigenomics. 1 Epi proColon Instructions for Use (IFU 0008) and Epigenomics data on file, P130001.Epigenomics Inc.
- Kato, K., Uchida, J., Kukita, Y., Kumagai, T., Nishino, K., Inoue, T., … Imamura, F. (2016). Transient appearance of circulating tumor DNA associated withde novotreatment.Scientific Reports,6, 38639. http://doi.org/10.1038/srep38639
- Loupakis, F., Ruzzo, A., Cremolini, C., Vincenzi, B., Salvatore, L., Santini, D., … Graziano, F. (2009).KRAScodon 61, 146 andBRAFmutations predict resistance to cetuximab plus irinotecan inKRAScodon 12 and 13 wild-type metastatic colorectal cancer.British Journal of Cancer,101(4), 715–721.http://doi.org/10.1038/sj.bjc.6605177
- Miller, K et al. Cancer Treatment and Survivorship Statistics, 2016. CA: A Cancer Journal for Clinicians 66: 271–289. doi:10.3322/caac.21349. Web. 28 Feb. 2018.
- Vashist, N. (2018). Liquid Biopsy: Revolutionizing Cancer Diagnoses and Treatments.BioChain.
- Vashist, N. (2018). A Tale of Two Biopsies: Liquid Biopsy vs Tissue Biopsy.BioChain. Warton, K., Mahon, K. L., & Samimi, G. (2016). Methylated circulating tumor DNA in blood: power in cancer prognosis and response. Endocrine-Related Cancer, 23(3), R157–R171. http://doi.org/10.1530/ERC-15-0369
Author
BioChain Institute Inc.

