What is RNA?
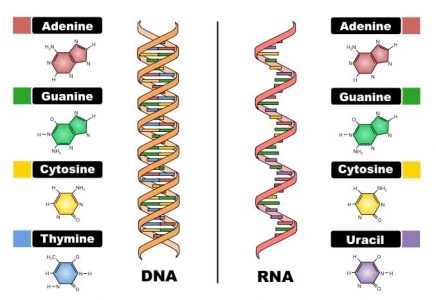
RNA (Ribonucleic acid) is found in the nucleus and is the result of the transcription of DNA (deoxyribonucleic acid) to RNA, which is then synthesized into protein. There are two main differences between DNA and RNA: 1) RNA contains three of the same bases as DNA guanine (G), adenine (A), and cytosine (C), but contains uracil (U) instead of thymine (T) and 2) RNA contains the sugar ribose instead of deoxyribose. Although the chemical differences between DNA and RNA are small, the structural differences can be significant. While DNA is always in a double helix, RNA is single-stranded and can therefore form an assortment of different shapes (Alberts et al.).
Why do we use RNA in Research?
Nearly every cell has the same set of genes. However, in each cell, only some of those genes are active and used to make proteins. Since each cell type can have different sets of genes “turned on”, researchers have begun investigating the downstream products of DNA such as RNA and protein. Studying the RNA and protein expressed in each cell allows the researcher to learn more about each individual cell type. Once RNA is extracted from a cell, it can inform us about which genes are being expressed and the relative abundance of those genes. However, shortly after researchers began studying RNA, they discovered that there are many different types of RNA, all of which play unique roles in the cell (Lodish et al.).
Types of RNA
There are several different types of RNA that have been identified to date. The three main types of RNA (mRNA, tRNA, and rRNA) work together to efficiently translate proteins. Each type of RNA has a distinct function and can be extracted from cells and studied experimentally.
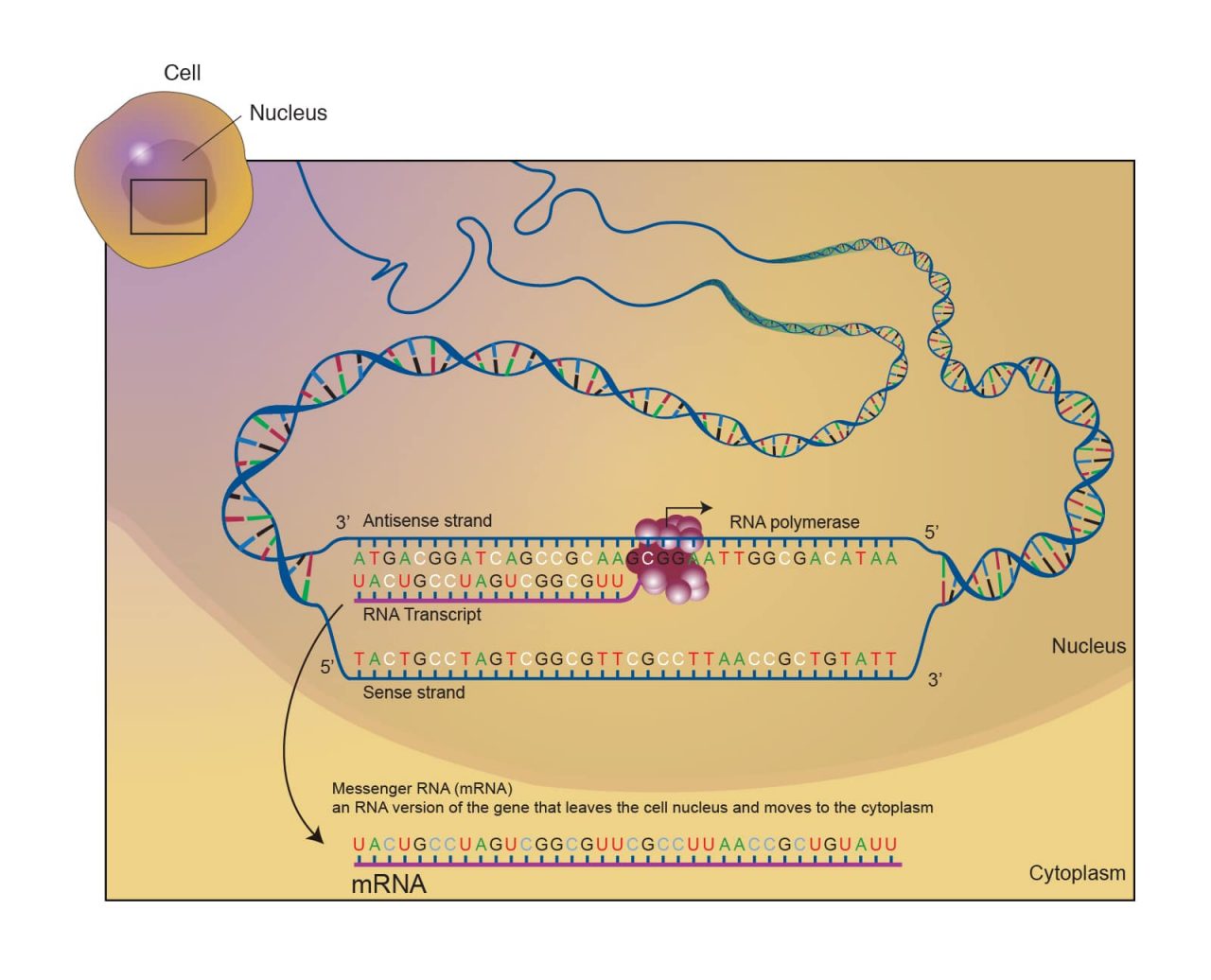
Messenger RNA (mRNA) mRNA is a copy of a portion of the DNA sequence that is first synthesized in cells, and then sent to the cytoplasm which can then determine the protein synthesized. The nucleotides in the mRNA sequence code for the order of amino acids (Lodish et al.). The quantity of mRNA for a certain gene can be determined by extracting total RNA, reverse transcribing the RNA into complementary DNA (cDNA) and then quantifying it using polymerase chain reaction (PCR) (Nolan et al.). If you are not familiar with mRNA extraction, you can also purchase normal and disease state mRNA to use in experiments from BioChain. (See Figure 1 Messenger RNA, mRNA)
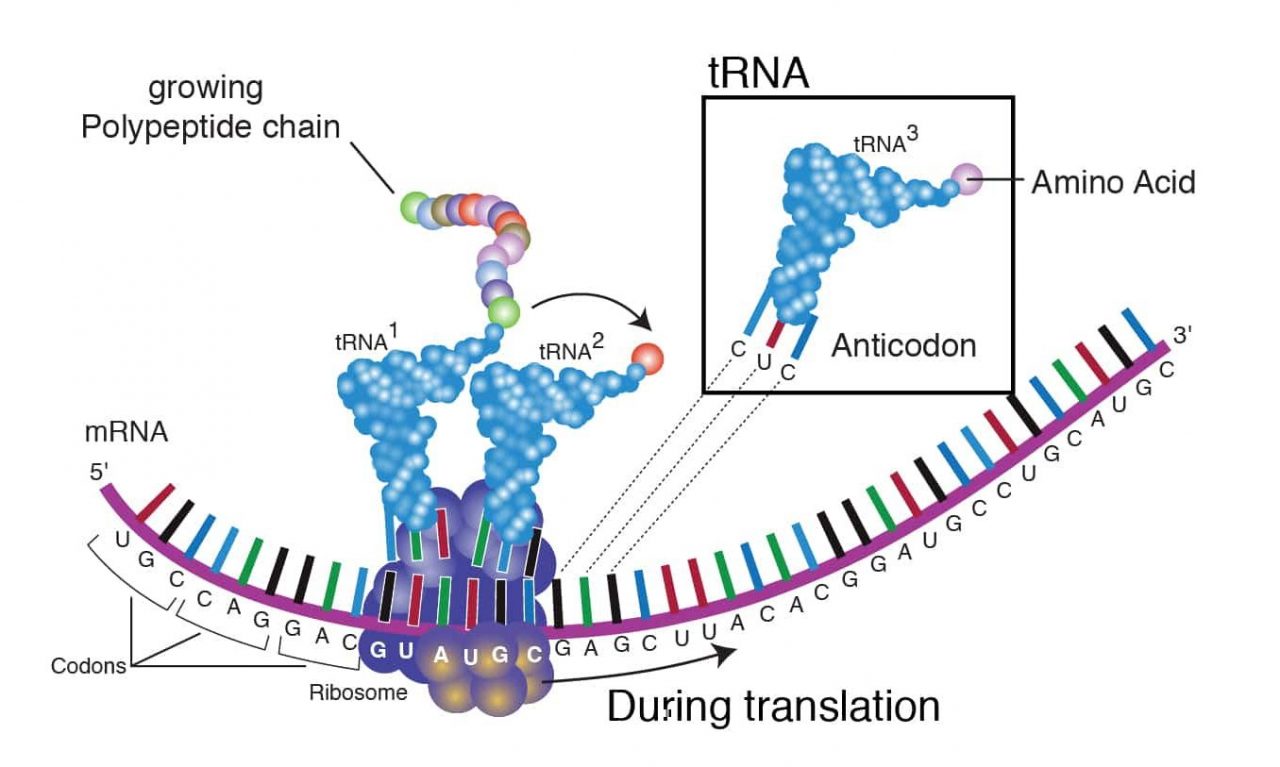
Transfer RNA (tRNA) tRNA is another type of RNA which “reads” the mRNA. During translation, each tRNA recognizes a certain three nucleotide mRNA sequence termed a “codon” and brings in the correct amino acid. The correct amino acid that is attached to the tRNA is then added to the growing amino acid chain, which will eventually become a functioning protein (Lodish et al.). tRNA can be extracted and studied using chromatographic and biochemical strategies. Recently, tRNA has been quantified using microarray assays (Kirchner et al.). (See Figure 2: Transfer RNA tRNA)
Ribosomal RNA (rRNA) rRNA is the essential component of the ribosome and assists in the translation of mRNA to protein. rRNAs have the most complex secondary structure of any RNA (Hyone-Myong Eun). The ribosomes move across mRNA and catalyze the reaction between the amino acids to form polypeptide chains and are also involved in binding tRNA and other proteins required to synthesize proteins. The ribosomes are composed of both a small and a large subunit, which work together to synthesize proteins (Lodish et al.).
Small interfering RNA (siRNA) The cell uses siRNA to interfere with and silence genes. siRNA is commonly used to downregulate certain genes to demonstrate a knockdown in cells or animal models. Two different complementary strands form the siRNA complex, one guide (antisense) strand and one passenger (sense) strand. Together they form a duplex, which is 19-25 bp with 3’dinucleotide overhangs. Synthetic siRNAs can be produced and transfected into cells or animals for knockdown experiments (Dharmacon and Dana et al.).
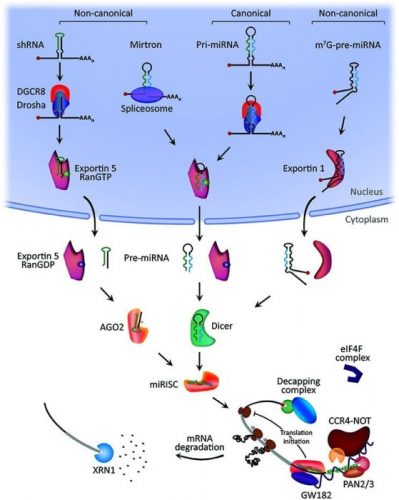
Micro RNA (miRNA) miRNAs are small non-coding RNA which are involved in the regulation of gene expression through mRNA. miRNA can achieve gene expression through many ways such as translational repression, mRNA cleavage, and deadenylation (Lodish et al.). miRNA research has grown over the past few years and miRNA can be isolated from total RNA and quantified using TaqMan analysis. miRNA can also be isolated from tissue culture cells, tissues, blood, and serum using miRNA isolation kits. (See Figure 3 MicroRNA biogenesis & Mechanism of Action)
Small nuclear (snRNA) snRNA’s are small non-coding RNA which are primarily located in the nucleus. snRNA’s are extremely important in splicing and form ribonucleoprotein (RNP). snRNAs can bind to certain sites in primary transcripts and help facilitate splicing (Valadkhan et al.).
Long non-coding RNA (lncRNA) Long non-coding RNA has recently become popular and is potentially a new form of regulation. There have been several lncRNAs identified, which have been connected to diseases. However, there is still much more to be discovered about this type of RNA (Kung et al.).
BioChain: RNA Isolation and Purchase
BioChain is proud to offer a range of kits to isolate total RNA as well as many other types of RNA. BioChain is also a leader in RNA collection and has an extremely large database of RNA from both disease states and normal tissues. BioChain offers total RNA from many different cell types and organs such as lymphocytes, cartilage, brain and many more.
BioChain also provides total RNA from different organisms, including humans (normal, diseased and tumor), mice, monkeys, rats and plants. They isolate the RNA thoroughly and make sure that there is no RNase, genomic DNA, proteoglycans and polysaccharide contamination in each sample. They check the 28s and 18s ribosomal RNA content to confirm that the integrity of the RNA is intact. BioChain also offers cDNA samples from various different disease states as well as normal tissues.
Whether you would like to isolate RNA from your own samples or receive RNA from matched normal and disease/tumor samples, BioChain can provide you with the materials necessary to achieve your research goal.
References
- Alberts B, Johnson A, Lewis J, et al. Molecular Biology of the Cell. 4th edition. New York: Garland Science; 2002. From DNA to RNA.Available from:
- Ressata. Introduction to Biological Principles. Pressbooks; 2013
- Breaker, R. R., Conklin, D. S., Gold, L., Söll, D., Montimurro, J. S., & Agris, P. F. (2012). RNA Science and its Applications—a look toward the future, RNA Biology, 9(8), 1050–1053, doi:10.4161/rna.21209
- Lodish H, Berk A, Kaiser C, et al. Molecular Cell Biology. 6th edition. New York: W.H. Freeman and Company; 2008.
- Nolan T, Hands RE, Bustin SA. (2006). Quantification of mRNA using real-time RT-PCR, Nature Protocols, 1(3): 1559-1582, doi:10.1038/nprot.2006.236
- Kirchner S., Rauscher R., Czech A., Ignatova Z., (2017). Microarray-Based Quantification of Cellular tRNAs, doi:10.17504/protocols.io.hfcb3iw
- Hyone-Myong Eun, (1996), Enzymology Primer for Recombinant DNA Technology, 1-108, doi:10.1016/B978-012243740-3/50004-1 http://dharmacon.horizondiscovery.com/applications/rna-interference/sirna/
- Dana H, Chalbatani GM, Mahmoodzadeh H, et al., (2017), Molecular Mechanisms and Biological functions of siRNA, 13(2):48-57 https://www.sigmaaldrich.com/life-science/functional-genomics-and-rnai/mirna/learning-center/mirna-introduction.html
- Valadkhan S. and Gunawardane LS, (2013), Role of small nuclear RNAs in eukaryotic gene expression, (54):79-90, doi:10.1042/BSE0540079
- Kung JTY, Colognori D and Lee JT, (2013), Long Noncoding RNAs: Past, Present, and Future, 193(3): 651-669, doi:10.1534/genetics.112.146704
- Yagi T., Kubota E., Koyama H., Tanaka T., Kataoka H., Imaeda K., Joh T., (2018), Glucagon promotes colon cancer cell growth via regulating AMPK and MAPK pathways, Oncogene
- https://www.biochain.com/search/?fwp_search_bar=RNA%20kits
- https://www.biochain.com/product/microrna-isolation-kit/
- Murata T. and Yamaguchi M., (2014), Alternatively spliced variants of the regucalcin gene in various human normal and tumor tissues, International Journal of Molecular Medicine: Spandidos Publications. https://www.spandidos-publications.com/ijmm/34/4/1141?text=fulltext
- Mahr A, Weinschenk T., Schoor O., Fritsche J., Singh H., Stevermann L., (2018), Novel Peptides and Combination of Peptides for use in Immunotherapy Against Various Tumors, United States Patent Application 20180030113, http://www.freepatentsonline.com/y2018/0030113.html
Author
BioChain Institute Inc.