An Introduction to PCR
Polymerase chain reaction (PCR) is a technique used to amplify a known focused segment of genomic DNA, RNA, or plasmid DNA to generate millions of copies of DNA/RNA segments that have multiple uses in the fields of molecular biology and other related fields. Genomic DNA is chromosomal DNA in organisms; in contrast, plasmids are circular DNA segments in bacteria that are independent of the chromosomal DNA. Majority of PCR methods involves exposure of the reactants to thermal cycler with repeated heating and cooling cycles under different temperatures to yield large number of DNA copies.
PCR Applications
- molecular biology for research
- DNA cloning and sequencing
- construction of DNA related phylogeny
- analysis of gene function
- diagnosis of hereditary diseases
- amplification of ancient DNA
- genetic fingerprinting for forensics analysis
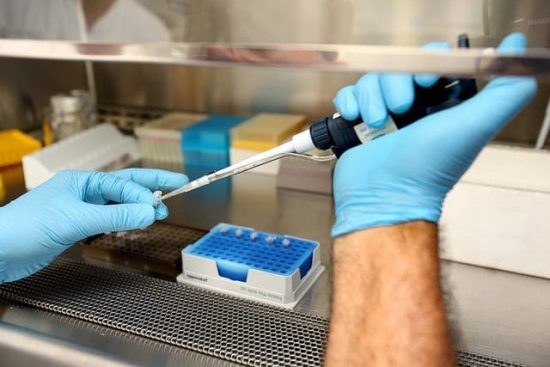
DNA Polymerase
All PCR applications employ heat stable DNA polymerase called Taq Polymerase, an enzyme isolated from thermophilic bacteria called thermus aquaticus. These bacteria live in hydrothermal vents and hot springs. The enzyme withstands protein denaturing conditions and is easy to use in PCR as it is subjected to varying degrees of extreme heat and cooling processes. Pfu Polymerase (isolated from pyrococcus furiosus) is a well-known enzyme which has lower error rates than Taq Polymerase but is very expensive. It can be used to get accurate PCR products.
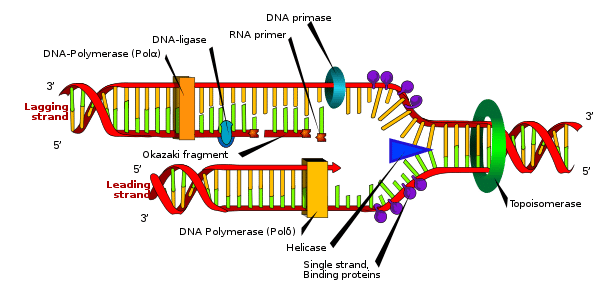
Figure: A representation of the structures of the replisome during DNA replication. Source
Oligonucleotide Primers
Primers are starting points for DNA synthesis. DNA polymerase can attach only to a double stranded DNA. DNA polymerase attaches to the primer and starts copying the DNA strand by adding the nucleotides. The procedure involves using two oligonucleotide DNA primers (short single stranded DNA sequences around 20-40 bases long) that are synthesized to complement the beginning and the ending sequence of the DNA strand to be copied. The primers have 40-60% GC rich content for optimal performance. Forward and reverse primers are used for amplification and they bind to the opposite strands of the DNA template by complementary base pairing. The separated DNA strands run in two directions: 3’ end to 5’ end on one, and 5’end to 3’end on the other; thus, two primers are used to amplify both strands.
PCR Procedure
The three stages of PCR are denaturation, annealing, and extension. There are seven core ingredients used in the setup of the procedure.
In a sterile microfuge tube the following reagents are mixed.
- Genomic DNA template (50ng - 250ng)
- Deionized water
- Reverse primer
- Forward primer
- DNA nucleotide bases(dNTPs)
- Taq DNA polymerase
- Taq assay buffer (contains Tris-HCl, MgCl2 and KCl).
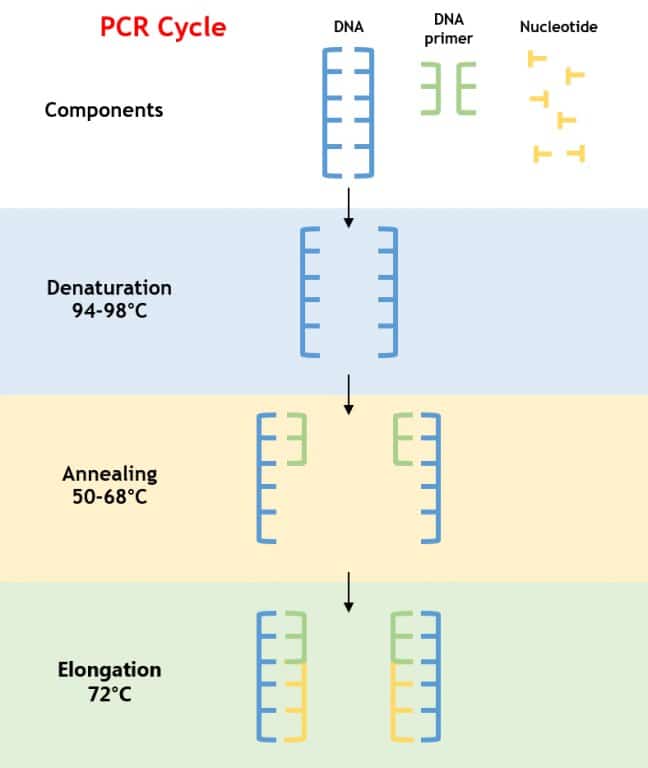
Denaturation
The above reagents are heated to 94°C for 3-5 minutes. (The high temperature causes the hydrogen bonds between the bases to break and the strands separate into two single DNA strands).
Annealing
The reaction is now cooled to 50-65°C for 10-30 seconds. (This allows the forward and reverse primers to attach to the single stranded DNA pieces through hydrogen bonds).
Extension
The reaction now is heated to 72°C for 5-10 minutes. (This is an optimal temperature for Taq Polymerase to attach to the primer and start adding the nucleotides one by one in 5’ to 3’ direction to build the complementary DNA strands).
The above steps are repeated 30 to 40 cycles to produce the sequence of interest. The process takes 1 minute to copy 1000 DNA bases. The whole process takes around 3 to 4 hours depending on the length of the fragment. The original DNA is not used as a template every time, as the new DNA made in one round can serve as a template for the next round of DNA amplification.
Visualization of PCR Results
Once PCR is completed, gel electrophoresis is used to check for quantity and size of the DNA fragments. Gel electrophoresis is a method where DNA fragments are dragged through a gel matrix using electric current to separate the DNA pieces according to size. A standard DNA ladder is included so the DNA sizes can be determined.

BioChain's PCR Kits: Ease and Convenience